BNFinder2 - new version available!
We have just released a new version of BNFinder. Since original
release in 2009, it contains a number of bugfixes and a whole new
set of functions. Most notable additions are:
- Support for classification with Bayesian networs including
cross-validation, ROC curve plotting and classification of new examples
- Inclusion of the Mutual Information Test (MIT) scoring
function for static/dynamic BNs with support for continuous variables
- Support for parallel computation on multiple CPU machines
- Possibility to pre-define the desired false-positive error-rate
during training
The new version is available for download from Python Package Index and
from our launchpad project
page.
You can also download documentation in pdf format as a single file or in separate parts:
Documentation sources and the exemplary input files are available in the source
package in the doc/ folder.
BNFinder - usage statistics
As a part of our efforts on releasing the new BNfinder version we have
looked at some statistics of BNfinder usage. Our input data was simply
the logs of our web interface (we cannot gather all download data from
different sites such as pypi and launchpad...).
All in all, within the three years sinc BNFinder got
published, our server served a bit more than 750 requests from a
bit over 220 users. We were curious where are our users from, so we
did a little analysis on the top-level domain parts of the registered
e-mail addresses. More than half (125) users have an e-mail address
somewhere on gmail.com or yahoo.com which prevents identification
but for the remaining users we mapped their location based on their
TLD (mapping .edu to US). The results are interesting:
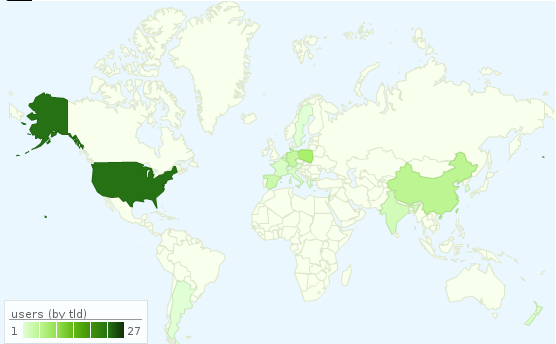
BNfinder seems to be used across almost-all continents (although it clearly needs
some publicity in South America and Africa ;)).
BNFinder - Bayesian Network Finder
BNF toolbox allows for Bayesian Network reconstruction. It can be used for both dynamic and static networks. It is written in python, and distributed under GNU GPL Library version 2.
The stable version of the software itself can be downloaded from here. A more recent, development version can be found on launchpad. It requires python version > 2.4,
(python 2.5 is recommended). Python 3.0 is not supported yet. Permanent installation is described in the installation istruction,
you can also run it without installing, just unpack the archive, go to the created directory and type:
python bnf --help
Documentation:
- Tutorial on using BNFinder in pdf file for printing,
html format or pdf in zip archive with data examples
- Description of methods
in html or pdf
- Users manual in html or pdf
- Whole documentation in a pdf version
If you do not want to install the software,
try our
Web interface. The interface is very simple (see
screenshot). In order to test it,
you only need to
provide an input file containing observed experimental data
(example file) and
your e-mail address. The resulting network (in a
SIF format readable by Cytoscape)
will be sent to your e-mail address.
If you have any questions or comments, please use our
mailinglist, or contact us directly at: [ dojer | bartek ]@mimuw.edu.pl,

